2017 Summer School
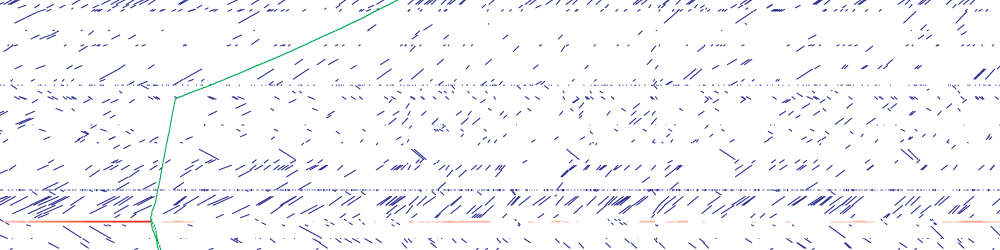
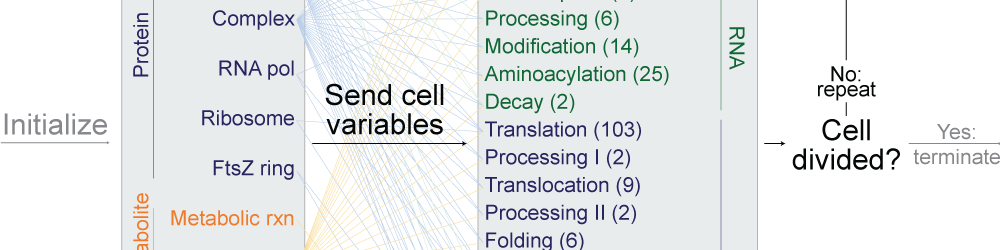


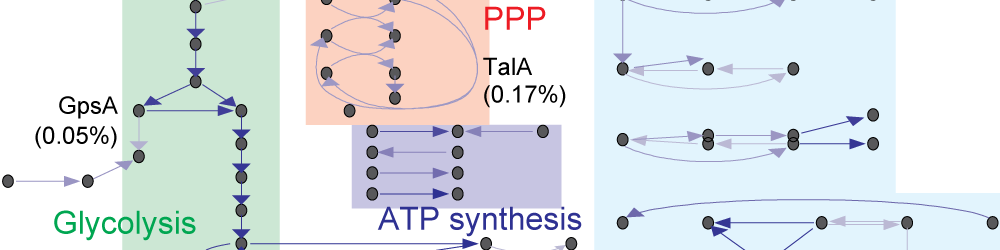

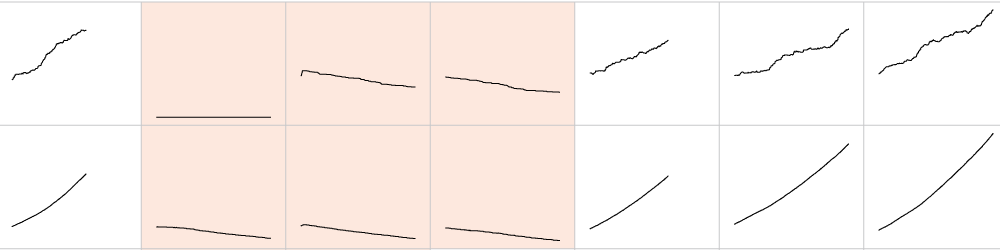
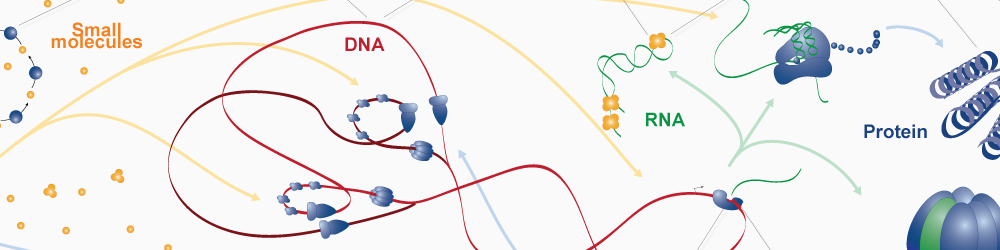
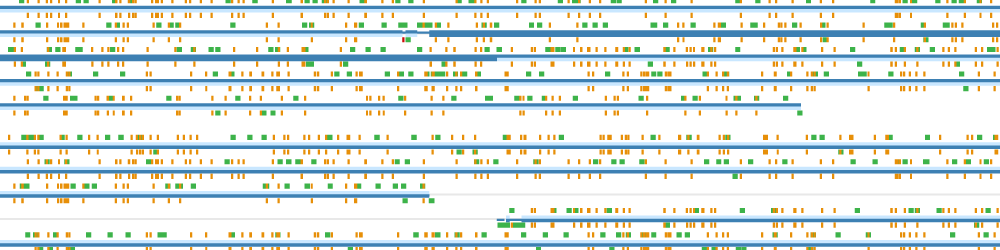
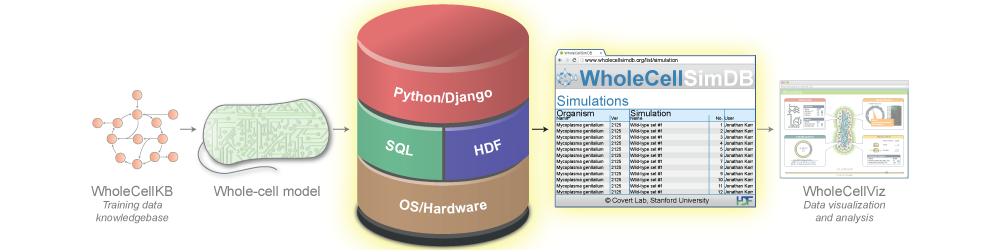
Whole-cell models are promising tools for predicting phenotype from genotype by accounting for every individual gene and cell function. Whole-cell modeling has the potential to enable rational bioengineering and precision medicine. However, significant work remains to develop fully complete and accurate whole-cell models. The goal of the 2017 Whole-Cell Modeling Summer School is to provide young investigators cutting-edge training in large-scale dynamical modeling and model integration.
Why participate?
The course will be the first course focused on multi-algorithm whole-cell modeling. It will teach strategies for building and managing large models which aren't covered by any other course including multi-algorithm modeling, model organism database curation, surrogate modeling, and software development. The five-day course will feature didactic lectures, interactive hands-on tutorials, and student research talks. The mornings will feature lectures on modeling individual pathways. The afternoons will feature interactive hands-on tutorials on building and analyzing multi-algorithm models to generate and evaluate hypotheses. Throughout the course, students will work toward building a small whole-cell model. In addition, the course will include student talks to enable students to share their own research.
Who is the course for?
The course is designed for PhD students and postdoctoral scholars who wish to gain training in large-scale dynamical modeling. See the pre-requesites section below for more information.
More info
Date & location
| Organizers
| Instructors
| Schedule
| Fees
| Scholarships
| Apply
| Directions
| Accomodations
| Course materials
| Photos
| Contact
Date and location
The summer school will be held September 4-7, 2017 at the Center for Regulatory Genomics in Barcelona, Spain.
Organizers
-

Jonathan Karr
Fellow, Icahn School of Medicine at Mount Sinai
Integrative modeling -

Maria Lluch-Senar
Staff Scientist, Center for Regulatory Genomics
Genomic profiling -

Luis Serrano
Group Leader and Director, Center for Regulatory Genomics
Genomic profiling
Instructors
-

Verónica Lloréns-Rico
PhD Student, Center for Regulatory Genomics -

Samuel Miravet-Verde
PhD Student, Center for Regulatory Genomics -

Saahith Pochiraju
Bioinformatician, Icahn School of Medicine at Mount Sinai -

Yosef Roth
Research Assistant, Icahn School of Medicine at Mount Sinai -

Balázs Szigeti
Postdoctoral Scholar, Icahn School of Medicine at Mount Sinai -

Marie Trussart
Postdoctoral Fellow, Walter and Eliza Hall Institute of Medical Research -

Marc Weber
Postdoctoral Fellow, Center for Regulatory Genomics
Content and schedule
The course will be four days long. The first day will feature an introductory lecture and ice-breaker activites. Day 2-4 will feature a combination of lectures, hands-on tutorials, and group discussions.
Pre-requisites
The course will focus on teaching students theory and techniques for large-scale dynamical modeling. Both computational and experimental researchers are encouraged to apply. However, due to the limited time of the course, the tutorials will assume prior knowledge of dynamical modeling (e.g. ordinary differential equations) and computer programming (e.g. Python). Participants who do not have experience with computer programming and/or dynamical modeling will be paired with participants who do to complete the tutorials. Similarly, participants who do not have extensive biological knowledge will be paired with participants who do.
Unfortunately, due to the limited time of the course, the tutorials will not have time to provide introductions to computer programming and dynamical modeling. There are several other courses which provide introductions to these topics:
- Coursera Systems Biology courses
- Advanced Lecture Course on Computational Systems Biology
- Dresden Summer School in Systems Biology
- In Silico Systems Biology
- qBio Summer School
| 9:00-10:30 am |
Tutorial
Modeling the 3-D structure of the chromosome Marie Tussart |
|---|---|
| 10:30-11:00 am | Coffee break |
| 11:00-12:30 pm |
Tutorial
Modeling the 3-D structure of the chromosome Marie Tussart |
| 12:30-2:00 pm | Lunch |
| 2:00-3:30 pm |
Tutorial
Pathway modeling using ordinary differential equations Verónica Lloréns-Rico |
| 3:30-4:00 pm | Coffee break |
| 4:00-5:30 pm |
Tutorial
Pathway modeling using ordinary differential equations Verónica Lloréns-Rico |
| 5:30-6:00 pm | Discussion |
| 9:00-10:30 am |
Tutorial
Genome-scale metabolic modeling using FBA Marc Weber |
|---|---|
| 10:30-11:00 am | Coffee break |
| 11:00-12:30 pm |
Tutorial
Genome-scale metabolic modeling using FBA Marc Weber |
| 12:30-2:00 pm | Lunch |
| 2:00-3:30 pm |
Tutorial
Stochastic simulation Jonathan Karr |
| 3:30-4:00 pm | Coffee break |
| 4:00-5:30 pm |
Tutorial
Stochastic simulation Jonathan Karr |
| 5:30-6:00 pm | Discussion |
| 9:00-10:30 am |
Tutorial
Model composition Jonathan Karr |
|---|---|
| 10:30-11:00 am | Coffee break |
| 11:00-12:30 pm |
Tutorial
Model composition Jonathan Karr |
| 12:30-2:00 pm | Lunch |
| 2:00-3:00 pm |
Tutorial
Rule-based modeling Samuel Miravet-Verde |
| 3:00-4:00 pm |
Tutorial
Data aggregation Yosef Roth and Saahith Pochiraju |
| 4:00-4:30 pm | Coffee break |
| 4:30-5:30 pm |
Tutorial
Model testing and validation Jonathan Karr |
| 5:30-6:00 pm |
Closing Maria Lluch-Senar and Jonathan Karr |
Recommended reading
Whole-cell modeling
- Karr JR, Takahasi K & Funahashi A. The principles of whole-cell modeling. Curr Opin Microbiol 27, 18–24 (2015).
- Carrera J & Covert MW. Why Build Whole-Cell Models?. Trends Cell Biol 25, 719–22 (2015).
- Macklin DN, Ruggero NA & Covert MW. The future of whole-cell modeling. Curr Opin Biotechnol 28, 111–115 (2014).
- Karr JR, Sanghvi JC, Macklin DN, Gutschow MV, Jacobs JM, Bolival B Jr, Assad-Garcia N, Glass JI & Covert MW. A whole-cell computational model predicts phenotype from genotype. Cell 150, 389–401 (2012).
- Tomita M, Hashimoto K, Takahashi K, Shimizu TS, Matsuzaki Y, Miyoshi F, Saito K, Tanida S, Yugi K, Venter JC et al. E-CELL: software environment for whole-cell simulation. Bioinformatics 15, 72–84 (1999).
Metabolic modeling
- Murabito E, Verma M, Bekker M, Bellomo D, Westerhoff HV, Teusink B & Steuer R. Monte-Carlo modeling of the central carbon metabolism of Lactococcus lactis: insights into metabolic regulation. PLoS One 9, e106453 (2014).
- Maarleveld TR, Khandelwal RA, Olivier BG, Teusink B & Bruggeman FJ. Basic concepts and principles of stoichiometric modeling of metabolic networks. Biotechnol J 8, 997–1008 (2013).
Intracellular signaling
- Klipp E & Liebermeister W. Mathematical modeling of intracellular signaling pathways. BMC Neurosci 7, S10 (2006).
Translation regulation
- Gorgoni B, Marshall E, McFarland MR, Romano MC & Stansfield I. Controlling translation elongation efficiency: tRNA regulation of ribosome flux on the mRNA. Biochem Soc Trans 42, 160–5 (2014).
Cell cycle and division regulation
- Di Ventura B, Knecht B, Andreas H, Godinez WJ, Fritsche M, Rohr K, Nickel W, Heermann DW & Sourjik V. Chromosome segregation by the Escherichia coli Min system. Mol Syst Biol 9, 686 (2013).
- Di Ventura B & Sourjik V. Self-organized partitioning of dynamically localized proteins in bacterial cell division. Mol Syst Biol 7, 457 (2011).
Integrative modeling
- Carrera J, Estrela R, Luo J, Rai N, Tsoukalas A & Tagkopoulos I. An integrative, multi-scale, genome-wide model reveals the phenotypic landscape of Escherichia coli. Mol Syst Biol 10, 735 (2014).
Logical modeling
- Morris MK, Saez-Rodriguez J, Sorger PK & Lauffenburger DA. Logic-based models for the analysis of cell signaling networks. Biochemistry 49, 3216–3224 (2010).
Rule-based modeling
- Sekar JA & Faeder JR. Rule-based modeling of signal transduction: a primer. Methods Mol Biol 880, 139-218 (2012).
Pathway/genome databases
- Keseler IM, Mackie A, Peralta-Gil M, Santos-Zavaleta A, Gama-Castro S, Bonavides-Martínez C, Fulcher C, Huerta AM, Kothari A, Krummenacker M, Latendresse M, Muñiz-Rascado L, Ong Q, Paley S, Schröder I, Shearer AG, Subhraveti P, Travers M, Weerasinghe D, Weiss V, Collado-Vides J, Gunsalus RP, Paulsen I & Karp PD. EcoCyc: fusing model organism databases with systems biology. Nucleic Acids Res 41, D605-12 (2013).
- Reed JL, Famili I, Thiele I & Palsson BO. Towards multidimensional genome annotation. Nat Rev Genet 7, 130-41 (2006).
Genomics
- Lluch-Senar M, Delgado J, Chen WH, Lloréns-Rico V, O'Reilly FJ, Wodke JA, Unal EB, Yus E, Martínez S, Nichols RJ, Ferrar T, Vivancos A, Schmeisky A, Stülke J, van Noort V, Gavin AC, Bork P & Serrano L. Defining a minimal cell: essentiality of small ORFs and ncRNAs in a genome-reduced bacterium. Mol Syst Biol 11, 780 (2015).
- Güell M, van Noort V, Yus E, Chen WH, Leigh-Bell J, Michalodimitrakis K, Yamada T, Arumugam M, Doerks T, Kühner S, Rode M, Suyama M, Schmidt S, Gavin AC, Bork P & Serrano L. Transcriptome complexity in a genome-reduced bacterium. Science 326, 1268–1271 (2009).
- Yus E, Maier T, Michalodimitrakis K, van Noort V, Yamada T, Chen WH, Wodke JA, Güell M, Martínez S, Bourgeois R, Kühner S, Raineri E, Letunic I, Kalinina OV, Rode M, Herrmann R, Gutiérrez-Gallego R, Russell RB, Gavin AC, Bork P & Serrano L. Impact of genome reduction on bacterial metabolism and its regulation. Science 326, 1263–1268 (2009).
- Kühner S, van Noort V, Betts MJ, Leo-Macias A, Batisse C, Rode M, Yamada T, Maier T, Bader S, Beltran-Alvarez P, Castaño Diez D, Chen WH, Devos D, Güell M, Norambuena T, Racke I, Rybin V, Schmidt A, Yus E, Aebersold R, Herrmann R, Böttcher B, Frangakis AS, Russell RB, Serrano L, Bork P & Gavin AC. Proteome organization in a genome-reduced bacterium. Science 326, 1235–1240 (2009).
Synthetic biology
- Gardner TS. Synthetic biology: from hype to impact. Trends Biotechnol 31, 123–5 (2013).
- Marchisio MA & Stelling J. Computational design tools for synthetic biology. Curr Opin Biotechnol 20, 479–85 (2009).
Registration fee
The registration fee includes all materials needed for the course, lunches, coffee breaks, a welcome dinner, and the city tour. Students are responsible for their lodging, breakfasts, and dinners.
- Academia: €400 ($430)
- Industry: €800 ($860)
Travel scholarships
Several scholarships will be available. We expect to be able to give several $1,000 to trainees from non-European institutions.
How to apply
The course application is available online . The application asks for brief descriptions of why you want to participate in the course, your background, and your current research, as well as your CV.
Selection criterion
Students will be selected based on the prior knowledge and research experience and their desire to participate in the course.
- Application posted: Spring, 2017
- Application due: May 18, 2017
- Notification of application decisions: June 2, 2017
- Summer school: September 4-7, 2017
How to get to the school @ The Center for Genomic Regulation (CRG)
The course will be held at the Center for Genomic Regulation (CRG) at the Parc de Recerca Biomedica de Barcelona (PRBB) at 88 Carrer del Doctor Aiguader, Barcelona 08003, Spain.
The CRG is accessible by taxi and bus/train from the Barcelona-El Prat Airport
- Taxi (15 min)
- Bus (1 h): (1) Take bus A1 toward Catalunya Place to Espanya Place. (2) Take bus D20 toward Passeig Maritim to Carrer Trewalny. (3) The PRBB is across the street.
Accommodations
Numerous hotels are available nearby (see Hotels.com map).
Course materials
All of the slides and course materials are available to the course participants at http://www.wholecell.org/school-2017/private. Please use the username and password provided by the course organizers.Photos
Contact and more information
Please contact Damjana Kastelic or Anna Sole Amat with any logistical questions. Please contact the organizers with any content questions.