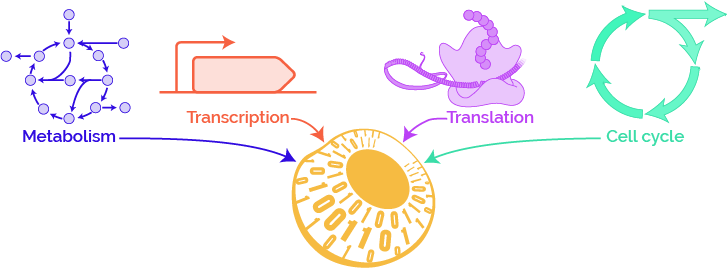
How will WC models predict phenotype from genotype?
Genome sequence
Genome
sequence
Genome sequence with single-nucleotide resolution
Molecular structures
Molecular
structures
Structure of each compound, chromosome, RNA transcript, and protein
Molecular interactions
Molecular
interactions
Participants and sites of each molecular interaction
Subcellular organization
Subcellular organization
Subcellular compartmentalization of each compound
Molecular concentrations
Molecular concentrations
Concentration of compound in each compartment
Thermodynamics and kinetics
Thermodynamics and kinetics
Equilibrium constant and rate law of each reaction
Stochastic variation
Stochastic
variation
Stochastic fluctuations of the concentration of each compound
Temporal dynamics
Temporal
dynamics
Temporal dynamics of each compound over the entire cell cycle
Spatial dynamics
Spatial
dynamics
Spatial distribution of each compound and reaction throughout cells
Population heterogeneity
Population heterogeneity
Stochastic intrinsic variation in the behavior of individual cells
Complex phenotypes
Complex
phenotypes
Complex phenotypes such as the growth rate and self-renewal
Extracellular environment
Extracellular environment
How cells behave in any extracellular environment